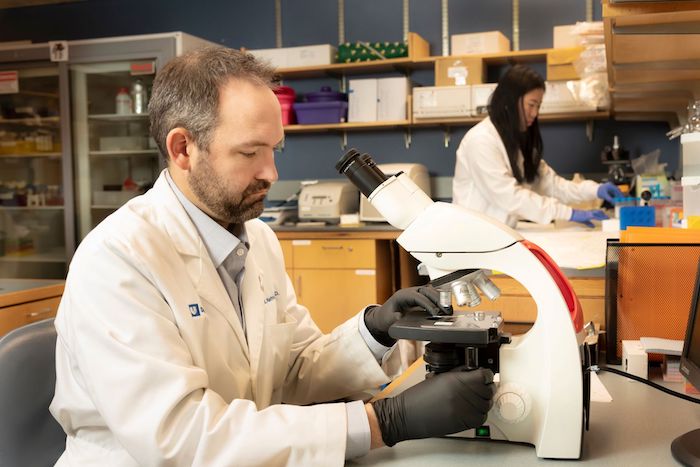
Reitman Lab Overview
Our laboratory studies how molecular alterations drive tumor heterogeneity and treatment response, with the goal of identifying new therapeutic vulnerabilities. We have a particular interest in pediatric and adult brain tumors. We develop genetically engineered mouse and human model systems to directly test how cancer-associated mutations shape tumor biology and therapy resistance. Using these platforms, we seek to uncover targetable mechanisms and design rational combination therapies, particularly at the intersection of radiation, molecular and immune therapies. We are especially interested in biological liabilities created by cancer mutations that can be exploited therapeutically. More broadly, we aim to leverage insights from cancer biology not only for treatment development but also for unconventional applications such as engineering useful proteins and biological tools inspired by disease mutations.
News
LaBella and Duke Faculty Among Authors of the 2024 BraTS-MEN-RT Dataset
Reitman Awarded ALSF Grant
FLASH Forward: Duke Researchers Explore Ultra-High Dose Rate FLASH Radiation Therapy for Brain Tumors
Vaios, Floyd, Reitman To Collaborate on Pilot Study
Reitman Lab Awarded 2024 DCI Grant
Reitman Awarded K08, New Investigator Award
Reitman Awarded Fellow Grant from The St. Baldrick’s Foundation
Team Member Spotlight: Zach Reitman, MD, PhD
Contact
Email Dr. Reitman at zjr@duke.edu.





